General Multimodal Protein Design Enables DNA-Encoding of Chemistry
May 9, 2026·,,,,,,,,,,,,,,,,,
Jarrid Rector-Brooks*
Théophile Lambert*
Marta Skreta*
Daniel Roth*
Yueming Long
Zi-Qi Li
Xi Zhang
Miruna Cretu
Francesca-Zhoufan Li
Tanvi Ganapathy
Emily Jin
Joey Bose
Jason Yang
Kirill Neklyudov
Yoshua Bengio
Alexander Tong
Frances H. Arnold
Cheng-Hao Liu
Abstract
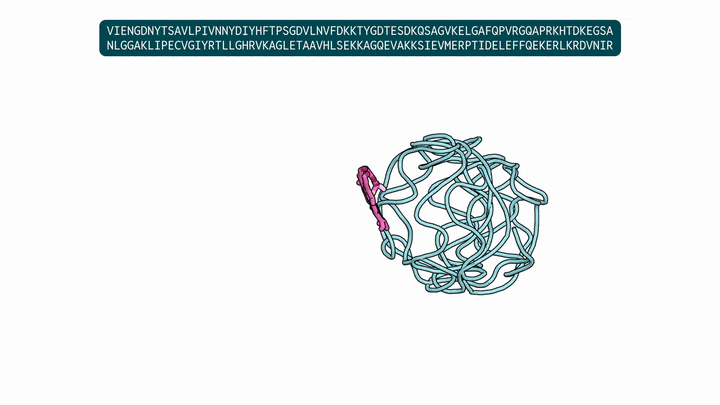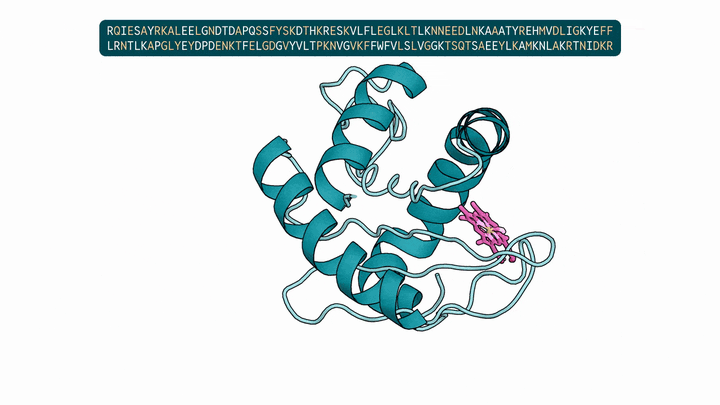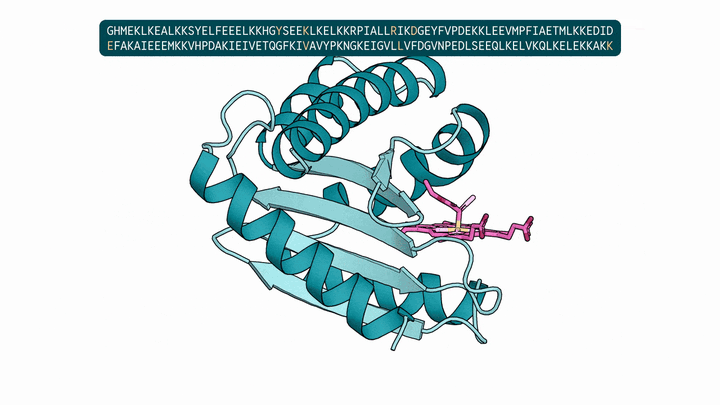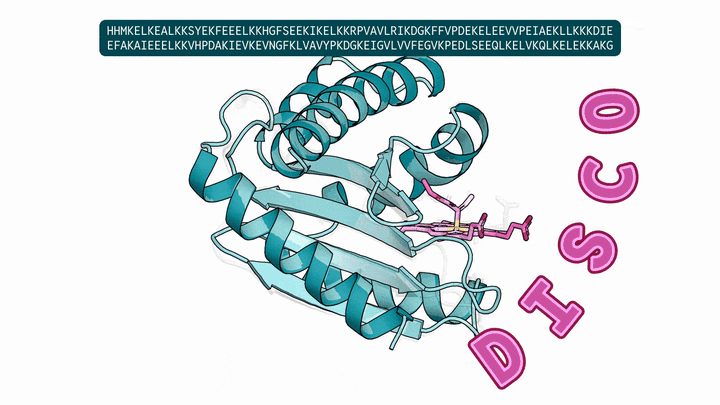
Evolution is an extraordinary engine for enzymatic diversity, yet the chemistry it has explored remains a narrow slice of what DNA can encode. Deep generative models can design new proteins that bind ligands, but none have created enzymes without pre-specifying catalytic residues. We introduce DISCO (DIffusion for Sequence-structure CO-design), a multimodal model that co-designs protein sequence and 3D structure around arbitrary biomolecules, as well as inference-time scaling methods that optimize objectives across both modalities. Conditioned solely on reactive intermediates, DISCO designs diverse heme enzymes with novel active-site geometries. These enzymes catalyze new-to-nature carbene-transfer reactions, including alkene cyclopropanation, spirocyclopropanation, B-H, and C(sp${}^3$)-H insertions, with high activities exceeding those of engineered enzymes. Random mutagenesis of a selected design further confirmed that enzyme activity can be improved through directed evolution. By providing a scalable route to evolvable enzymes, DISCO broadens the potential scope of genetically encodable transformations.
Type
Publication
aRxiv